
from QC to gene prediction and phylogenomics
We are pleased to announce the release of new BUSCO datasets! Based on OrthoDBv12.2 (https://orthodb.org), the new datasets represent a significant increase in coverage over all domains. For eukaryota, the new odb12.2 datasets represent a significant increase in coverage, especially in mammals, along with a more rigorous and balanced sampling of species. For prokaryota, the odb12.2 datasets are the same as the odb12 datasets released in 2025.
BUSCO v6.1.0 is the current stable version!
Gitlab, a Conda package and Docker container are also available.
Based on evolutionarily-informed expectations of gene content of near-universal single-copy orthologs, the BUSCO metric is complementary to technical metrics like N50.
Getting started — BUSCO tutorial
A concise walkthrough of BUSCO usage and workflows. Click to watch on YouTube →
Cite us
The latest BUSCO paper, describing the BUSCO v6 and the new OrthoDB.v12 BUSCO datasets is here, the correct citation would be:
Tegenfeldt F., Kuznetsov D., Manni M., Berkeley M., Zdobnov E.M., Kriventseva E.V. OrthoDB and BUSCO update: annotation of orthologs with wider sampling of genomes. Nucleic Acids Research, Volume 53, Issue D1, 6 January 2025, Pages D516–D522, https://doi.org/10.1093/nar/gkae987
The previous BUSCO paper, describing the BUSCO v4 and v5 and the OrthoDB.v10 datasets.
The following protocol covers the various BUSCO running modes and workflows, BUSCO setup, guidelines to interpret the results, and additional analyses, e.g., for building phylogenomic trees and visualizing syntenies using BUSCO results.
*Manni M., Berkeley M. R., Seppey M., & Zdobnov E. M. (2021).
License
The BUSCO software is licensed under the MIT License.
The BUSCO datasets are licensed under the Creative Commons Attribution-NoDerivatives 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nd/4.0/ or send a letter to Creative Commons, PO Box 1866, Mountain View, CA 94042, USA.
Any use of these datasets for analyses in a publication or product must include the citation of the corresponding paper:
- https://doi.org/10.1093/nar/gkae987 for _odb12* datasets, or
- https://doi.org/10.1093/molbev/msab199 for _odb10* datasets.
Obtain BUSCO
Full installation instructions are provided in the user guide and protocols.
BUSCO is available as a conda package and as a Docker image. Both of these versions are ready to run out of the box. Alternatively, it is also possible to manually install BUSCO.
The BUSCO software directly downloads the necessary datasets, whether they are specified by the user or automatically selected by BUSCO.
To display all available datasets
busco --list-datasets
You can also download them manually.
Earlier versions: v4, v3, v2, v1
Documentation and support
User guide Issues board"Core" genes
Mandatory Options
-i # Input sequence file or folder
-m # BUSCO analysis mode to run. Can be 'genome', 'protein' or 'transcriptome'.
Recommended Options
-c # Number of threads/cores to use
-l # Specify the BUSCO lineage dataset to be used for scoring
-o # Specify the name of the output folder
Pipeline-specific options
Augustus pipeline
--augustus # Invoke the BLAST/Augustus pipeline
-e # E-value cutoff for BLAST searches.
--limit # How many BLAST candidate regions to consider per BUSCO (default: 3)
--long # Optimization Augustus self-training mode (Default: Off)
--augustus_parameters # "--PARAM1=VALUE1,--PARAM2=VALUE2"
--augustus_species AUGUSTUS_SPECIES # Specify a species for Augustus training
Metaeuk pipeline
--metaeuk # Invoke the Metaeuk pipeline
--metaeuk_parameters # "--PARAM1=VALUE1,--PARAM2=VALUE2"
--metaeuk_rerun_parameters # "--PARAM1=VALUE1,--PARAM2=VALUE2"
Miniprot pipeline
--miniprot # Invoke the miniprot pipeline (default for eukaryota, option for prokaryota)
Genome Mode (all)
--skip_bbtools # Skip BBTools for assembly statistics
--scaffold_composition # Writes ACGTN content per scaffold to a file scaffold_composition.txt
--contig_break n #Number of contiguous Ns to signify a break between contigs. Default is n=10.
Other options
--config CONFIG_FILE # Provide a config file
--download [dataset ...] # Can be a dataset name, "all", "prokaryota", "eukaryota" or "virus"
--download_path DOWNLOAD_PATH # Set a custom local location for downloaded files
-f # Force overwrite an output directory
-r # Restart a previous incomplete run
--list-datasets # List all available BUSCO datasets
--offline # Indicate BUSCO should not attempt to download files
-q # Quiet mode
--tar # Compress some output subdirectories




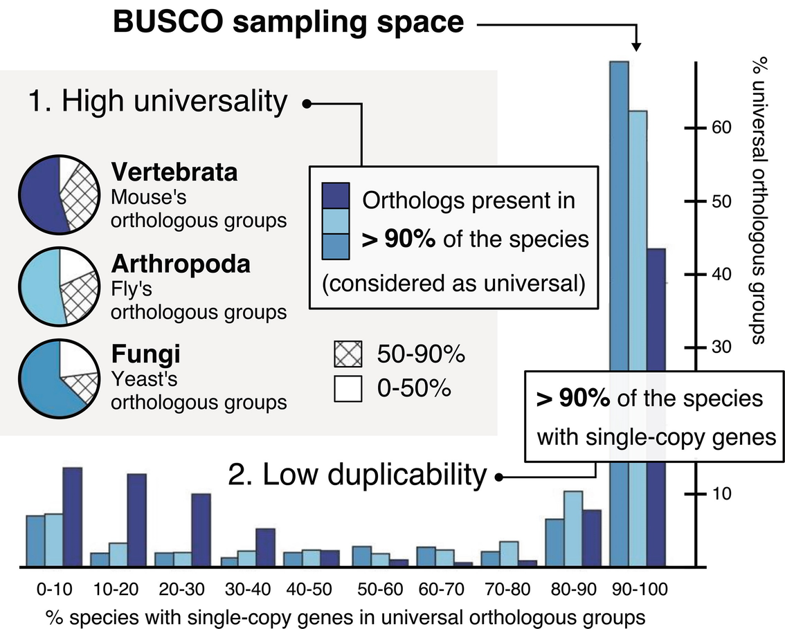 adapted from
adapted from